top of page

FOCUSED ON PATIENTS
DRIVEN BY DISCOVERY
Base4 is redefining RNA-targeted therapeutics through novel small molecule modulation to go after previously untreatable diseases. Powered by our revolutionary structure-based discovery platform, we are solving complicated diseases in several disease areas including neurology, oncology, and rare diseases.
...and others
Anchor 1

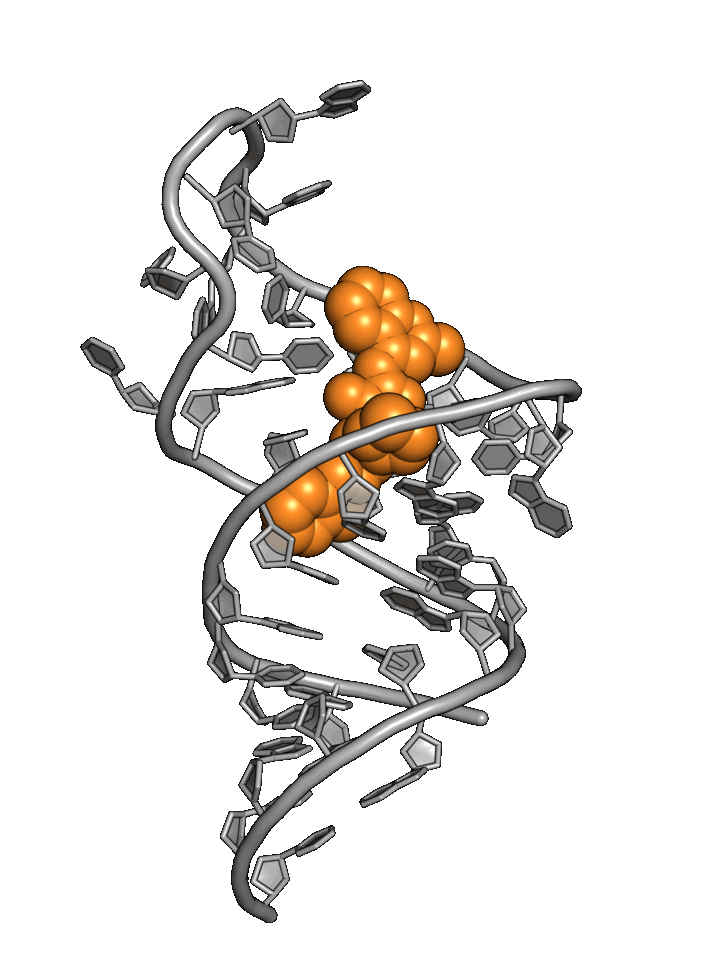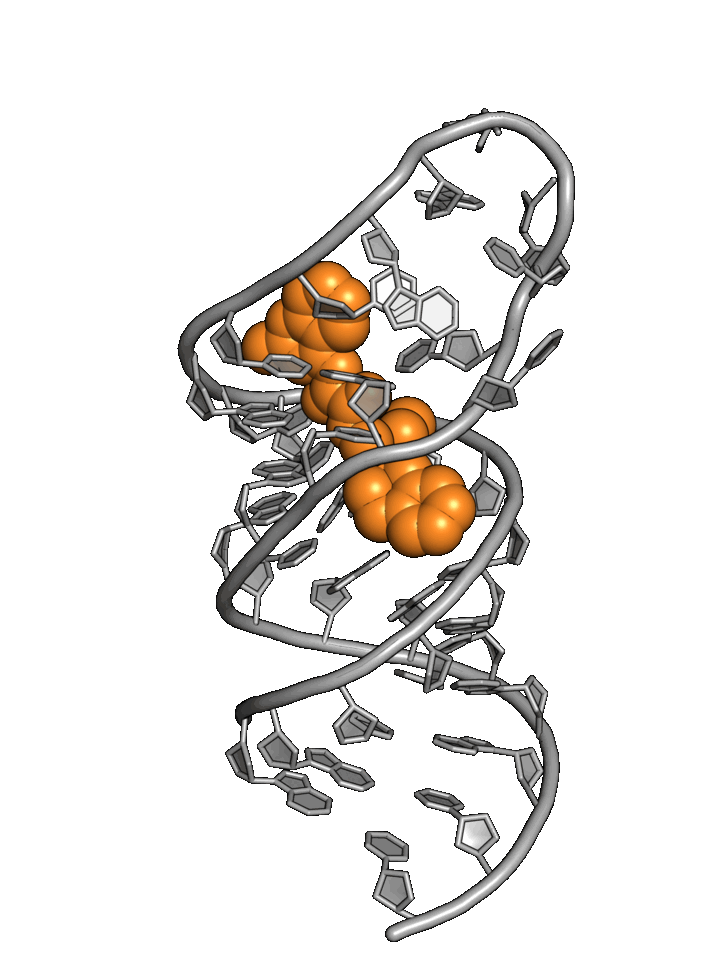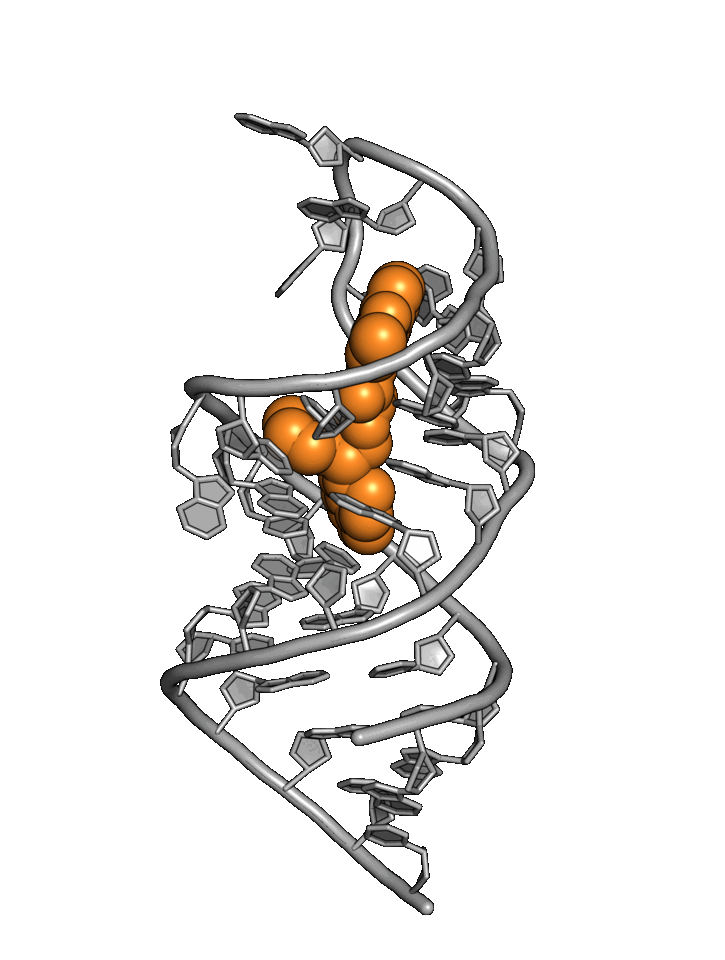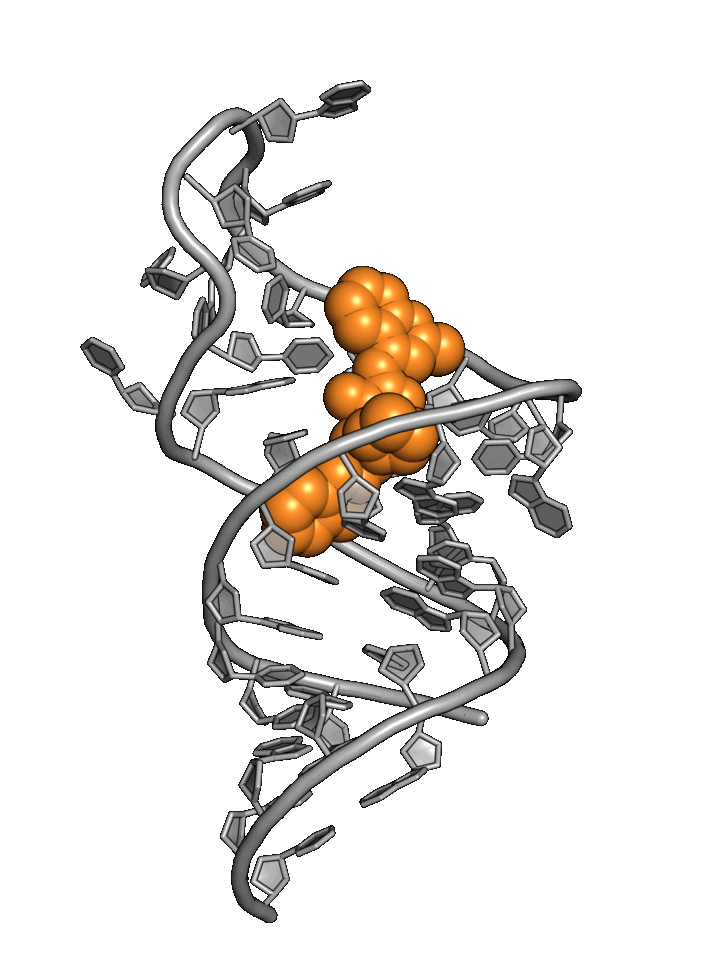
Exploiting RNA
structural dynamics
RNA structure wiggles and jiggles, forcing drug hunters to rely primarily on phenotypic methods
Base4 addresses RNA targets with a best-in-world structure-based approach.
We leverage our proprietary DART Platform, an AI-enhanced, quantum biophysical discovery platform that resolves dynamic atomic-resolution structures of RNA targets and screens for selective molecules against druggable RNA regions


bottom of page